Research
[fourcol_one]
[/fourcol_one] [fourcol_three_last]
Introduction
Blood vessels supply the nutrients necessary for organ & tissue function—every action is dependent upon a healthy, intact vasculature. Indeed, over 70 diseases are angiogenesis-related. Cancers represent a well-known angiogenesis-related disease. Here, abnormal blood vessels sustain tumor growth, development, and metastasis. However, anti-angiogenic therapies only moderately affect patient survival, ultimately leading to patient resistance.
We believe that unraveling vascular signaling complexities can occur by engineering quantitative experimental tools and computational models. Our research directly addresses this need, applying a “bottom-up,” systems biology paradigm to measure, integrate, and simulate mechanisms regulating angiogenesis.
[/fourcol_three_last]
[hr]
[fourcol_one]
[/fourcol_one]
[fourcol_three_last]
Quantitative Data Needed for Computational Models
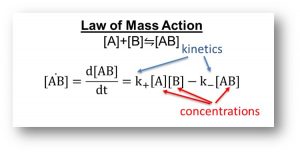
Our computational models are governed by mass action kinetics, which are defined by both the amount of the species (concentrations), and the probability of these species interacting (kinetics). Furthermore, our mass action kinetic models specifically examine protein signaling. So the data that we need to parameterize our models are both protein concentrations and protein-protein interaction kinetics.
While there are a plethora of data available on protein expression (e.g., Western blots) and protein-protein interactions (e.g., Co-IP) much of these data, report relative protein amounts or qualitative descriptions of interactions. Unfortunately, there are few quantitative data available. Our work aims to overcome these challenges by measuring both plasma membrane protein concentrations and protein-protein interaction kinetics.
[/fourcol_three_last]
[hr]
[fourcol_one]
[/fourcol_one] [fourcol_three_last]
Making Protein Presence Quantitative
In addition to parameterizing computational models, many receptors distinguish differences in cells, or cell “heterogeneity.” Indeed, tumors are often classified by the presence or absence of receptors (e.g., breast cancer classification: triple negative, ER-/PR-/Her2-). Therefore, our receptor quantification adds objectivity to tumor classification—providing absolute numbers that set standards for diagnosis. Such standards can be compared across samples and research groups, while addressing the significant biomedical challenge of understanding cell and tumor heterogeneity.
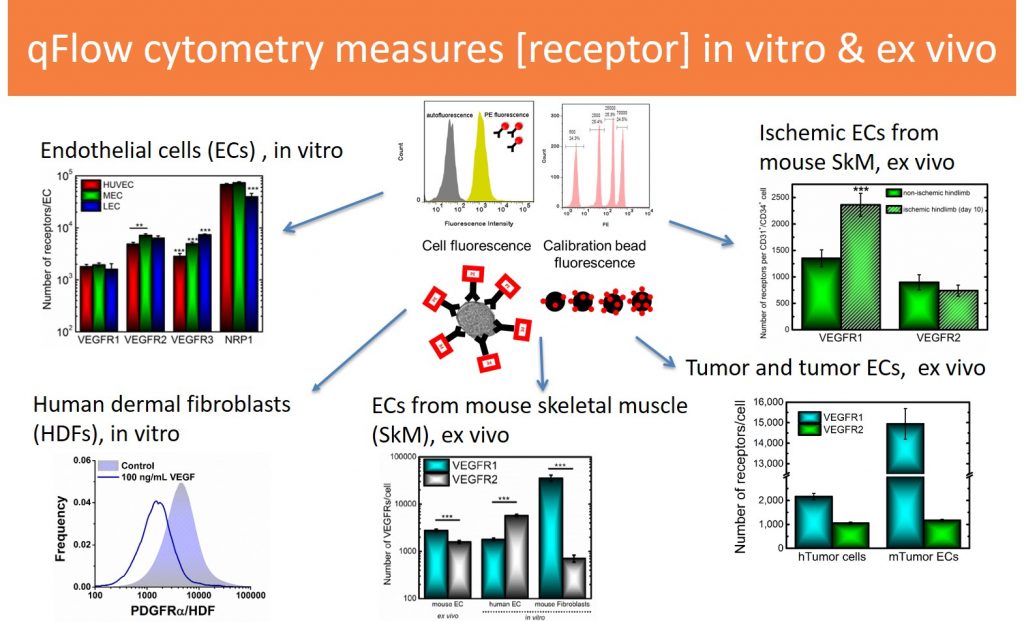
Towards these goals of parameterizing computational models, standardizing biomedicine, and understanding heterogeneity, we advance methods for quantifying plasma membrane concentrations of several receptors, including VEGFRs, neuropilins (NRPs), PDGFRs, EGFRs, Angiopoietin receptors, and others. Our primary method is quantitative flow cytometry. We are currently expanding our approach to enable multiplexed receptor quantification.
In vitro receptor quantification: We have applied qFlow cytometry to measure protein concentrations on several commonly used primary and expanded cell lines, in vitro, including: HUVECs, human dermal microvascular endothelial cells (HDMECs), human dermal lymphatic microvascular endothelial cells (HDLMECs), human dermal fibroblasts (HDFs), human tumor cells (MDA-MB-231, MCF-7, and U87), mouse fibroblasts (BALB/3T3 clone A31), mouse macrophages (RAW 264.7). We are currently expanding our approach to measure receptors on cells from peripheral blood samples.
Ex vivo receptor quantification: We have quantified receptors on endothelial cells from mouse skeletal muscle and two pre-clinical models of human vascular dysfunction: peripheral artery disease (PAD) and breast cancer. We are currently applying our approach to quantify receptors in glioblastomas.
[/fourcol_three_last]
[hr]
[fourcol_one]
[/fourcol_one]
[fourcol_three_last]
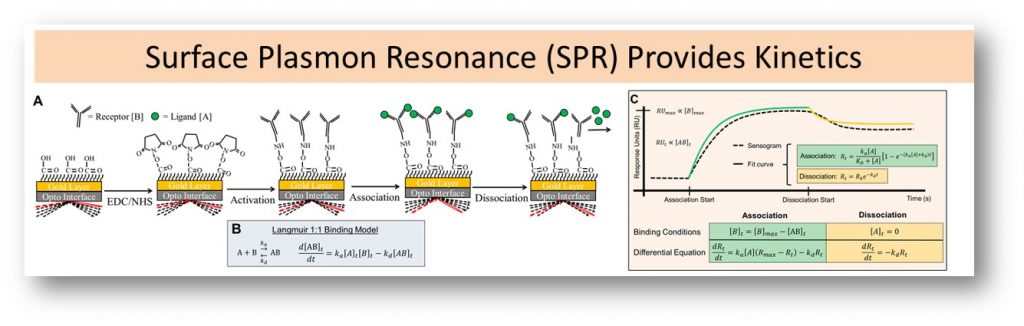
Exploring a New Signaling Paradigm: Cross-family Binding
Our computational models require quantitative data on protein-protein interaction kinetics. We apply a surface plasmon resonance approach to measure these interactions. Through our work, we have discovered high-affinity cross-family binding (e.g., PDGF–VEGFR). We are currently expanding our measurements of cross-family binding, and measuring signaling to understand its implications.
[/fourcol_three_last]
[hr]
[fourcol_one]

[/fourcol_one] [fourcol_three_last]
New Biological Insight via Computation
Our experimental data are providing measurements of receptor concentrations and they have discovered new ligand-receptor interactions that may change the way we interpret signaling. We have integrated these data into computational models and arrived at several key insgihts. Our computational models have integrated pre-clinical VEGFR1 concentrations to predict that high-concentrations of VEGFR1 may may negate anti-VEGF therapy. We have computationally examined how the structure of the VEGFR1 leads to cell signaling. We are exploring how cross-family binding affects signaling. Furthermore, with a cross-family binding paradigm, we can imagine the possibility of “ligand-promiscuity.” Wherein, several cross-family binding combinations are possible, each with differing signaling sensitivity. We are taking the first steps towards understanding combinatorial signaling: ranking the amount of signaling occurring via the canonical pathways (e.g., PDGFRs, IGFR1, EGFR, VEGFRs, Tie2, and FGFR1). Together this integrative approach is advancing us towards rational control of angiogenesis.
[/fourcol_three_last]
[hr]
[fourcol_one]
[/fourcol_one]
[hr]