Disease resistance in maize
Disease resistance in maize
Plants are in constant contact with microbes that utilize diverse mechanisms of attack. To protect themselves plants have evolved layers of defense that encompass diverse mechanisms of resistance and are shaped by the evolutionary dynamics between the plant and the microbe. The Jamann lab focuses on host-microbe interactions in maize and sorghum. The goals of the program are to mine genetic variation for resistance, determine how those genes are influencing the interaction, and better understand how pathosystems co-evolve. We study a diversity of pathogens and are taking a multi-faceted approach utilizing genetics, genomics, molecular biology, and evolutionary biology to understand the interaction between host and microbe. Students and professionals explore questions related to host-microbe interactions and develop a variety of skills related to these approaches. Ultimately exploiting genetic variation and understanding the governing mechanisms will lead to the development of more resistant varieties.
Specific projects:
Conserved genetic mechanisms for biotic stress in sorghum
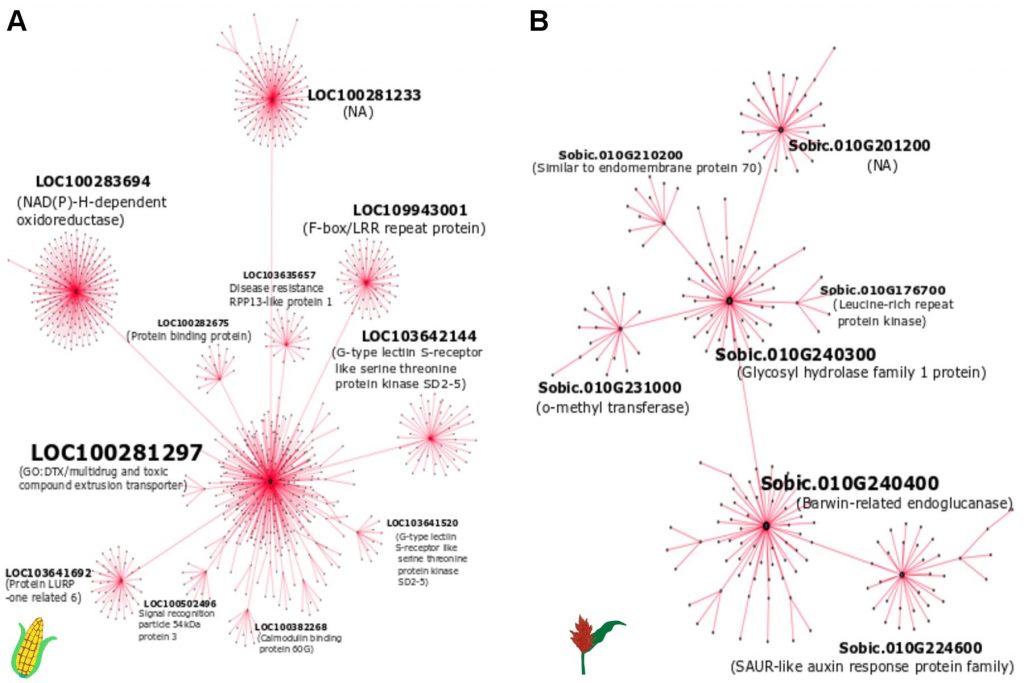
Setosphaeria turcica (syn. Exserohilum turcicum) can infect both maize and sorghum, yet isolates are host specific. Sorghum leaf blight (SLB), caused by S.turcica, is widespread and can decrease yields, reduce forage quantity and quality, and predispose plants to other diseases such as anthracnose (causal organism: Colletotrichum sublineolum). Host resistance is one of the most environmentally friendly and cost-effective methods of disease control, and resistance can be conserved across different plant species. By leveraging the knowledge of resistance in maize, we will accelerate the improvement of resistance to S. turcica in sorghum, while also developing this as a system to understand how microbes evolve to become pathogens of bioenergy crops. Furthermore, genes can confer resistance to other pathogens when tested in new systems, and we will assess the relationship between resistance to sorghum leaf blight and anthracnose. Our overall objective is to improve biotic stress resistance in sorghum, and to achieve this we will use a paired strategy of identifying plant genes that will confer resistance and identifying fungal genes that are key deterrents of a host jump from corn to sorghum. The identification of the genes involved in this interaction will enhance the prospects for strategic deployment of sustainable host resistance-based approaches in bioenergy crops.
Funding: Department of Energy, Plant Feedstock Genomics
Dissection of bacterial disease resistance in maize
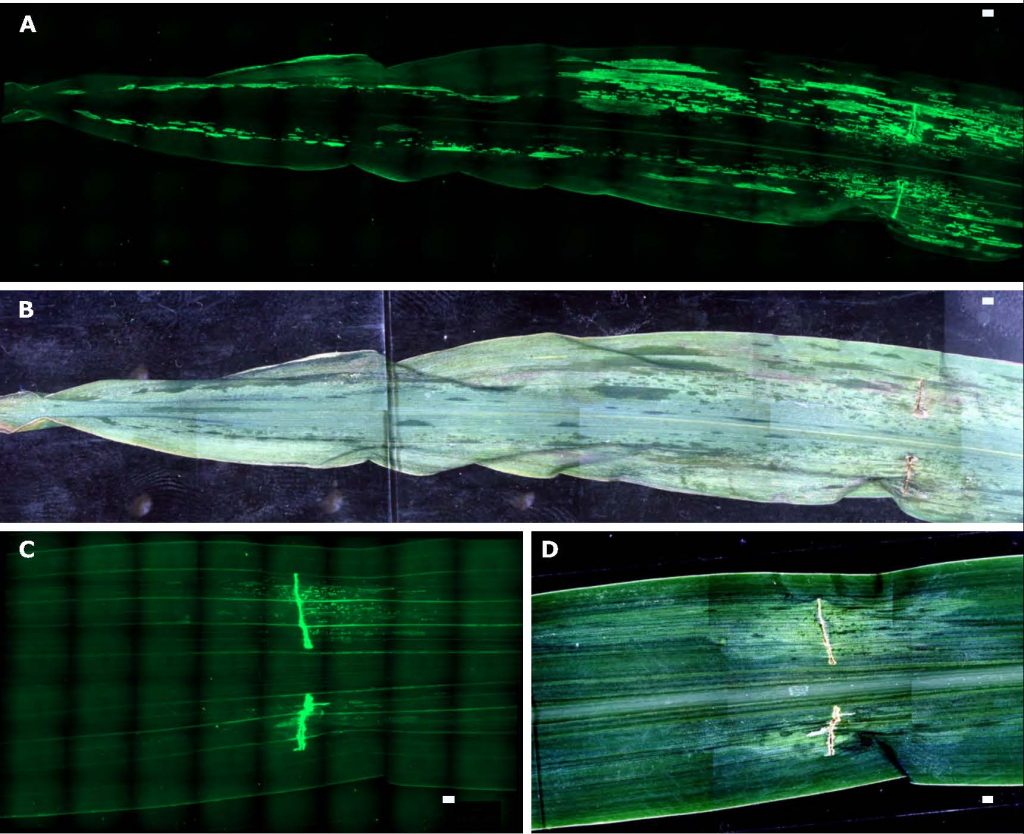
There are several significant bacterial diseases of maize, including Goss’s bacterial wilt (caused by Clavibacter nebraskensis) and blight and bacterial leaf streak (caused by Xanthomonas vasicola pv. vasculorum). We have mapped resistance for both diseases and identified regions that confer resistance to multiple diseases of maize (Qiu et al. 2020 G3; Qiu et al. 2020 Crop Science; Cooper et al. 2018 Crop Science). While identifying regions conferring resistance is critical to breeding more resistant varieties, understanding the host-pathogen interaction is imperative for deploying resistance in a strategic manner. For Goss’s bacterial wilt and blight we developed a system to visualize infection in planta and examined colonization and movement patterns of Clavibacter nebraskensis during infection using green fluorescent protein (GFP)-labeled bacterial strains (Mullens and Jamann 2021 Plant Disease). Resistant maize lines exhibited decreased bacterial spread in the vasculature and the mesophyll. We continue to examine both the genetic and mechanistic basis of bacterial disease resistance in maize.
Funding: Foundation for Food and Agricultural Research, Rapid Outcomes for Agricultural Research