We are interested in the fine details of cellular processes and the large cellular machines that perform them. We are best known for revealing structures and functions of the ribosome, RNAs and RNA-binding proteins involved in the cellular translation process that is fundamental to gene expression and regulation. Our mission is to identify the molecular mechanisms of these essential biological processes using a combination of biochemical, biophysical, structural, and molecular biology techniques.
Our Vision
Regulation of translation and quality control directly influences cellular processes such as development, differentiation, growth, cell fitness and cellular adaptive responses to the external environment. Regulatory dysfunction underpins various human diseases including metabolic disorders, neuronal degenerative diseases and cancer. We hope to gain knowledge of how dysregulation is associated with human diseases and to provide insights into possible new treatment strategies in medicine.
“All truths are easy to understand once they are discovered; the point is to discover them.” – Galileo Galilei
Molecular Mechanisms of Translational Regulation
We aim to define how regulation of translation alters critical aspects of cellular function that are essential for life. The unexpected gap between the expression of the genome at the level of transcriptome and proteome demonstrates the pervasiveness and importance of translational regulation. Regulation at the translational level not only provides a critical and fine-tuned control in gene expression, but also offers the cell an opportunity to rapidly respond to internal cellular cues and external stimuli without invoking nuclear events that involve transcription, mRNA processing and transport. We are interested in a range of regulatory mechanisms including exploring functions of the 5’-untranslated region (5’UTR), cap-independent translation and nonAUG translation. In practice, these works frequently involve creating or adopting new methodologies, analyzing data sets generated at the genomic level, and solving atomic structures of macromolecules.
Molecular Mechanisms of Quality Control
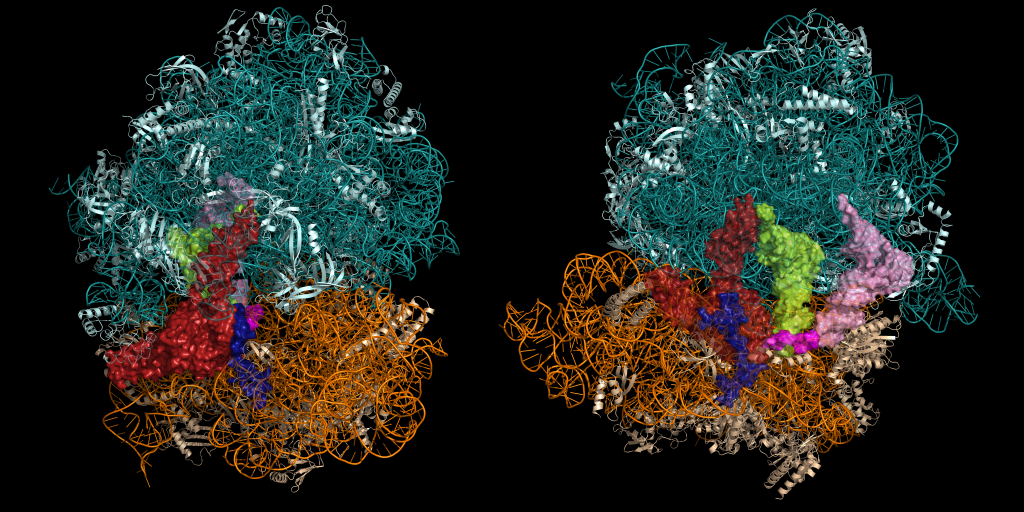
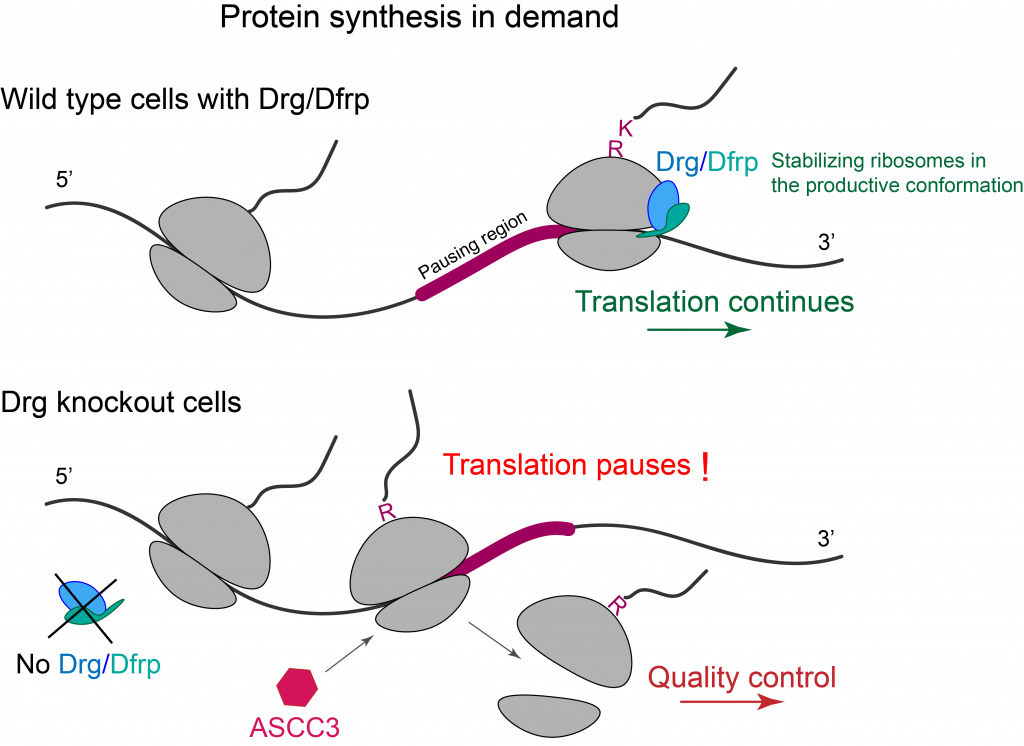
Cells have evolved elegant and sophisticated surveillance mechanisms to ensure protein homeostasis. The ribosome has been shown to be a central hub in sensing changing environments and regulating stress responses and gene expression. In this project, we study the molecular functions of highly conserved, developmentally regulated GTP-binding (Drg) proteins and their interactions with ribosomes and microtubules. Drg proteins are highly expressed in actively growing and developing cells, as well as reproductive adult tissues of plants, animals, and humans. Consistent with their functions in growth control, cell differentiation, and proliferation, Drg proteins are directly linked with human cancers, and developmental and neurological disorders. We discovered, for the first time, that Drg proteins alleviate translation pause and promote peptide-bond formation by stabilizing the paused ribosome in a productive conformation. We are currently investigating molecular mechanisms underpinning this important cellular process.
Functions of RNA-binding proteins in the post-transcriptional regulation
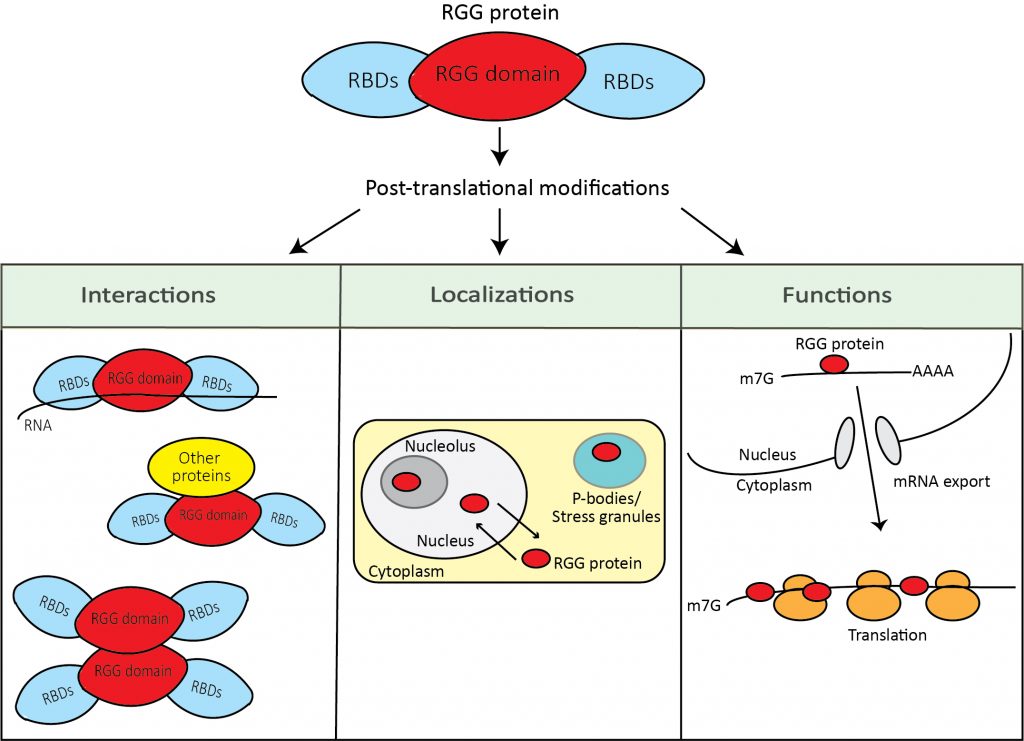
Thousands of proteins in the human genome contain RGG or RG repeats, and they are functionally involved in nearly every cellular pathway. Versatile, specific yet adaptable molecular interactions enabled by the unique combination of arginine and glycine, RGG motif proteins interact with a broad range of proteins and nucleic acids. More proteins containing functional RGG repeats, and their biological targets are bound to be uncovered in the future. Furthermore, post-translational modifications at the arginine in the motif extend the RGG protein’s capacity for a fine-tuned regulation. We study the properties, abundance, interactions, subcellular localization, and structures of RGG proteins in the cell. We aim to understand the mechanisms by which interactions in ordered or disordered domains, short motifs, and post-translational modification states synergistically function to control the protein functions.
Noncoding RNA functions in gene expression
Noncoding RNAs play essential and diverse roles in regulating gene expression. For example, base pairings between small nucleolar RNAs (snoRNAs) and sequences in eukaryotic rRNAs target specific nucleotides for modifications. rRNA modifications are critical for ribosome biogenesis, assembly, structure, and translating function, and most of the modifications are clustered in the functionally important regions of the ribosome. snoRNAs play other essential functions in the cell such as participating in pre-mRNA processing, directing alternative splicing, and acting as microRNA precursors. We study how different snoRNAs influence ribosome functions and dynamics.
Bacterial ribosome rescue as a new antimicrobial strategy
Bacteria use a unique trans-translation from eukaryotes to rescue stalled ribosomes, a process that can be exploited to develop new antibiotics in response to pressing clinical challenges of antibiotic resistance. Intervening after an error occurs is essential in maintaining protein homeostasis and contributing to the overall accuracy and fidelity of translation in living cells. In this process, cellular signals that initiate the rescue are first deciphered in the ribosome. Using bacterial proteins such as ArfA/RF2 as the model system, we determined structures, functions, and mechanisms important to the rescue process. We establish that the kinetics of ArfA/RF2-mediated rescue of bacterial ribosomes follows an induced-fit mechanism and illustrate the essential molecular interactions responsible for its mechanism of action. Results from this basic science research have been further used to develop new antibiotics for bacterial infection.